Provides a simple way to generate plots of pdfs, probability mass functions, cdfs, probability histograms, and normal-quantile plots for distributions known to R.
plotDist(
dist,
...,
xlim = NULL,
ylim = NULL,
add,
under = FALSE,
packets = NULL,
rows = NULL,
columns = NULL,
kind = c("density", "cdf", "qq", "histogram"),
xlab = "",
ylab = "",
breaks = NULL,
type,
resolution = 5000L,
params = NULL
)Arguments
- dist
A string identifying the distribution. This should work with any distribution that has associated functions beginning with 'd', 'p', and 'q' (e.g,
dnorm(),pnorm(), andqnorm()).distshould match the name of the distribution with the initial 'd', 'p', or 'q' removed.- ...
other arguments passed along to lattice graphing routines
- xlim
a numeric vector of length 2 or
NULL, in which case the central 99.8 of the distribution is used.- ylim
a numeric vector of length 2 or
NULL, in which case a heuristic is used to avoid chasing asymptotes in distributions like the F distributions with 1 numerator degree of freedom.- add
a logical indicating whether the plot should be added to the previous lattice plot. If missing, it will be set to match
under.- under
a logical indicating whether adding should be done in a layer under or over the existing layers when
add = TRUE.- packets, rows, columns
specification of which panels will be added to when
addisTRUE. SeelatticeExtra::layer().- kind
one of "density", "cdf", "qq", or "histogram" (or prefix of any of these)
- xlab, ylab
as per other lattice functions
- breaks
a vector of break points for bins of histograms, as in
histogram()- type
passed along to various lattice graphing functions
- resolution
number of points to sample when generating the plots
- params
a list containing parameters for the distribution. If
NULL(the default), this list is created from elements of\dotsthat are either unnamed or have names among the formals of the appropriate distribution function. See the examples.
Details
plotDist() determines whether the distribution
is continuous or discrete by seeing if all the sampled quantiles are
unique. A discrete random variable with many possible values could
fool this algorithm and be considered continuous.
The plots are done referencing a data frame with variables
x and y giving points on the graph of the
pdf, pmf, or cdf for the distribution. This can be useful in conjunction
with the groups argument. See the examples.
See also
Examples
plotDist('norm')
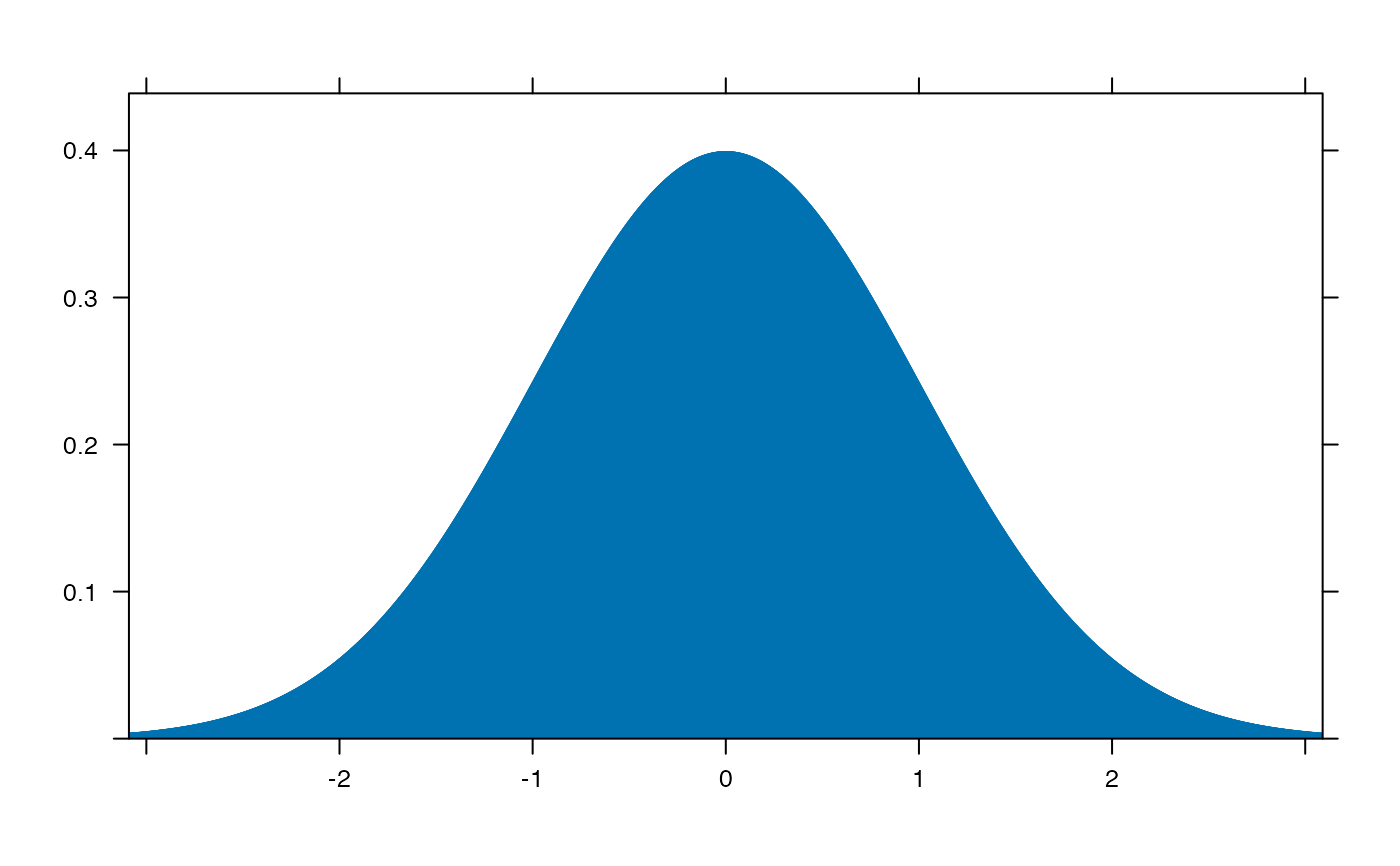
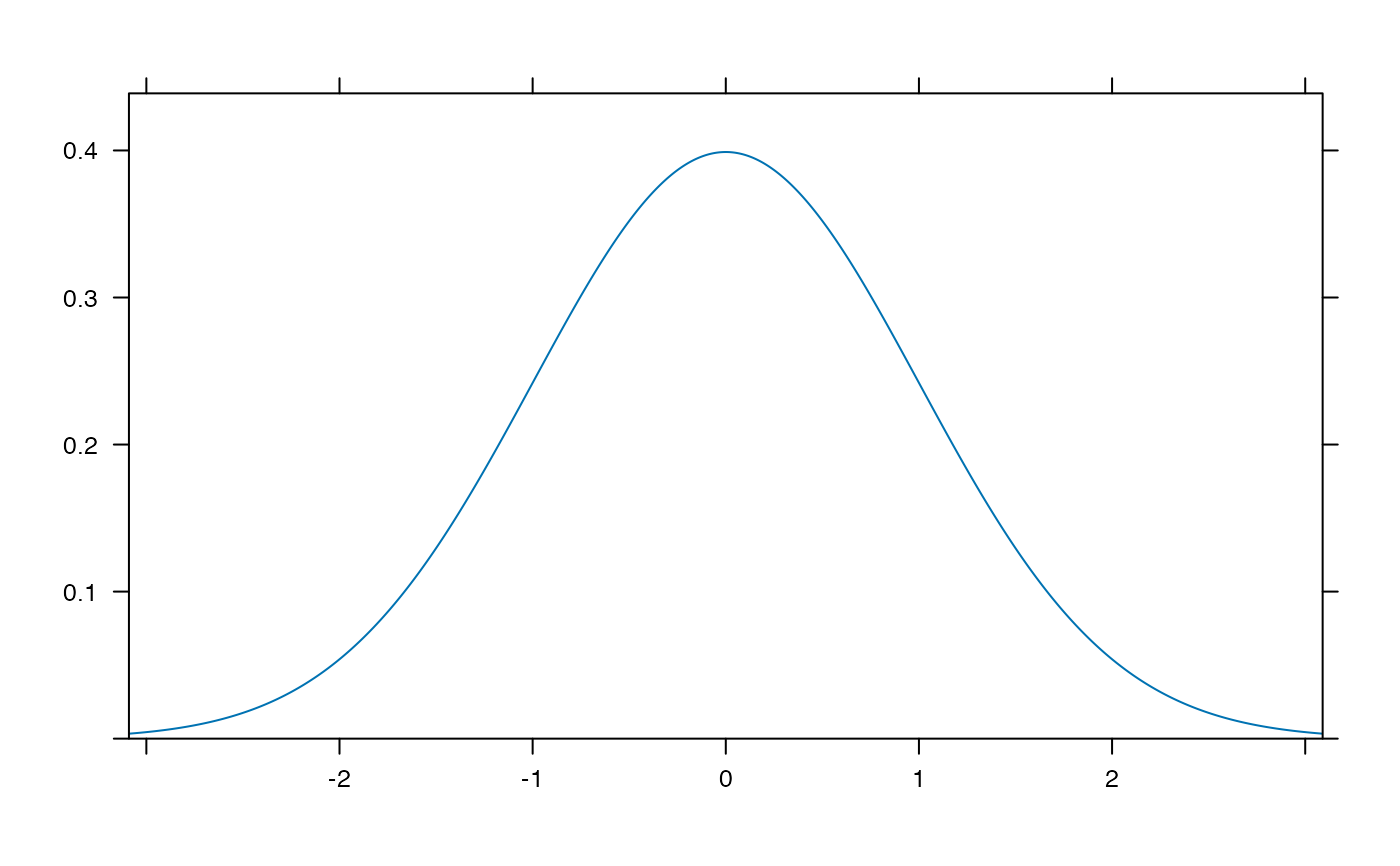 plotDist('norm', type='h')
plotDist('norm', type='h')
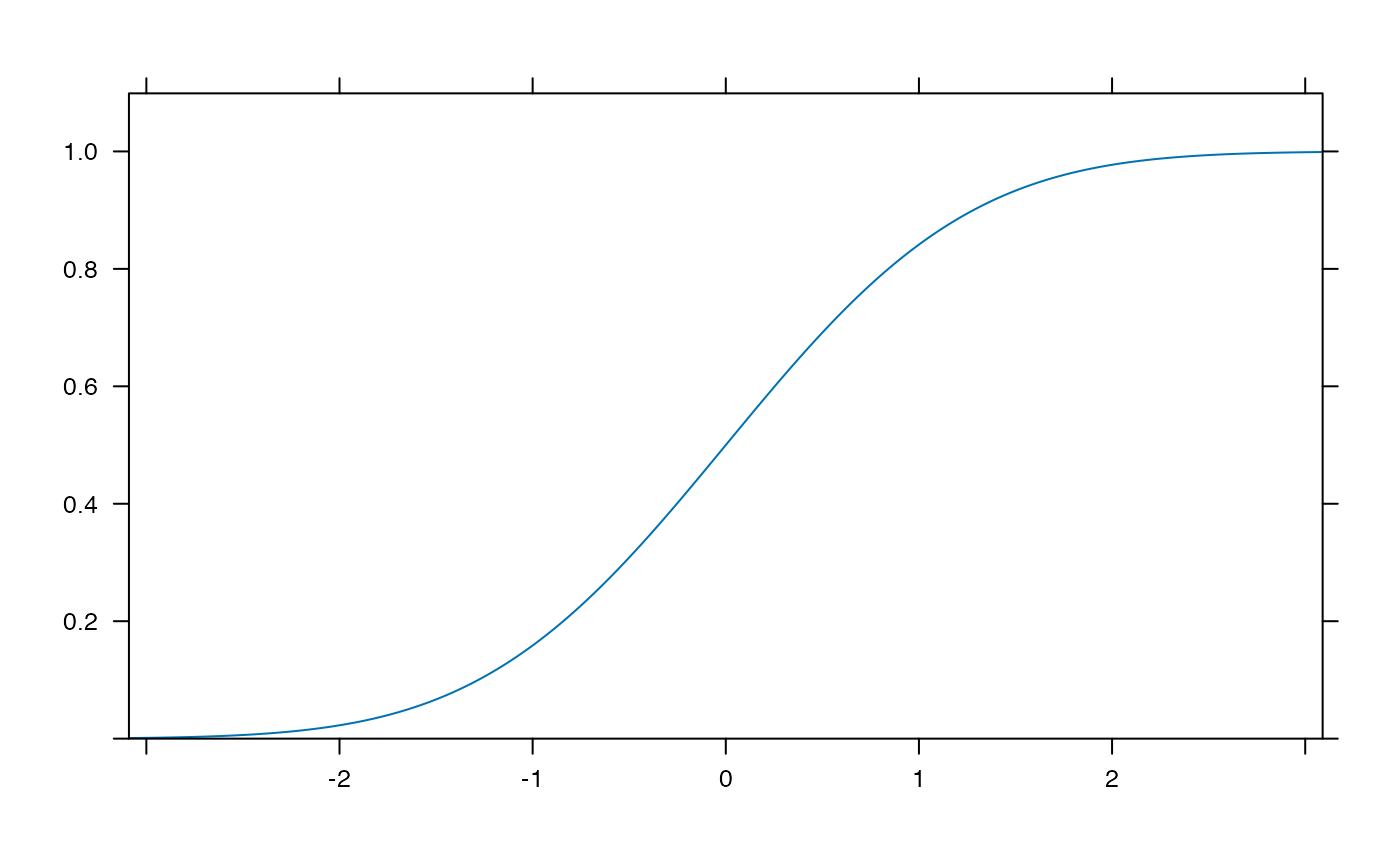
 plotDist('norm', kind='cdf')
plotDist('norm', kind='cdf')
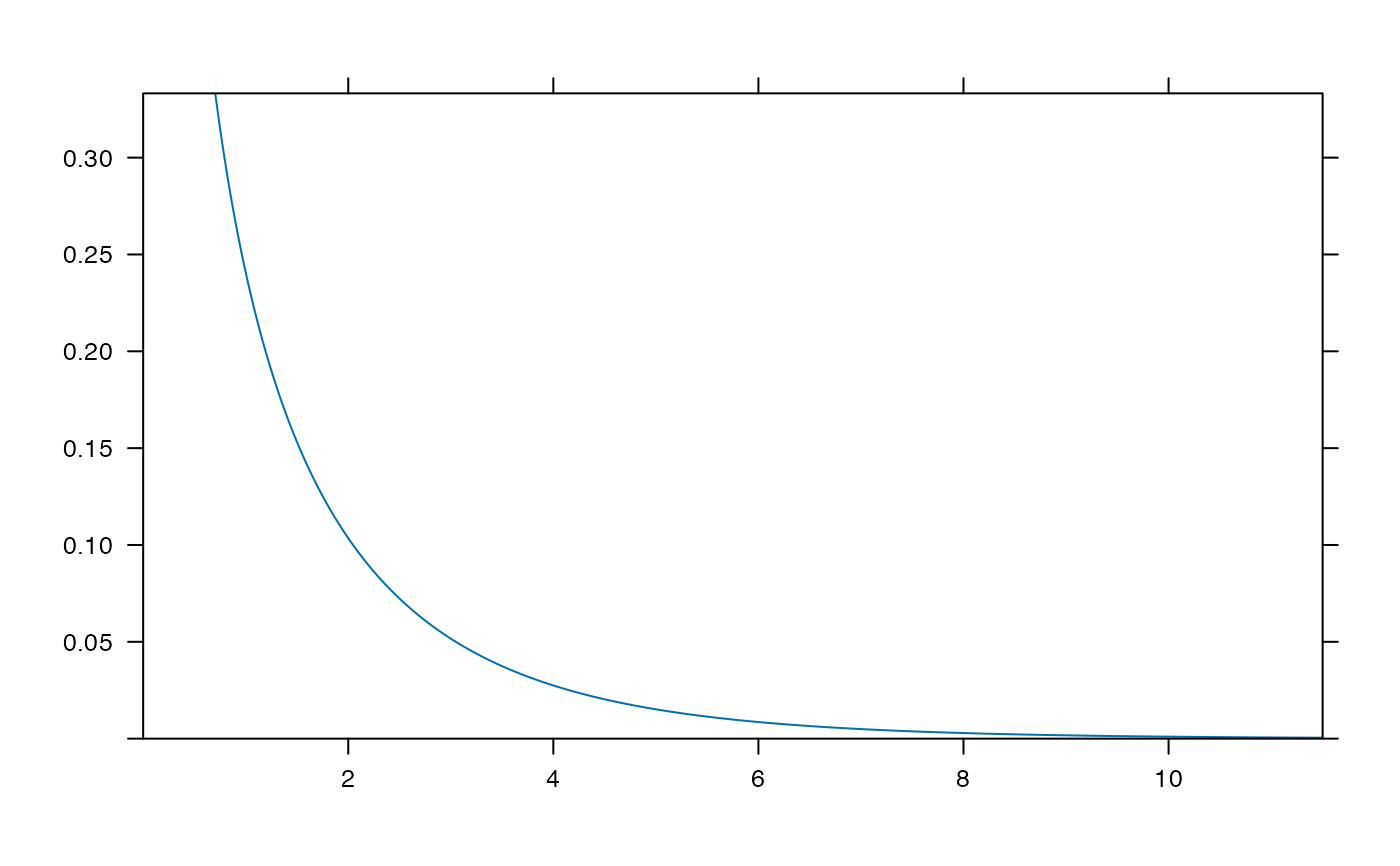
 plotDist('exp', kind='histogram')
plotDist('exp', kind='histogram')
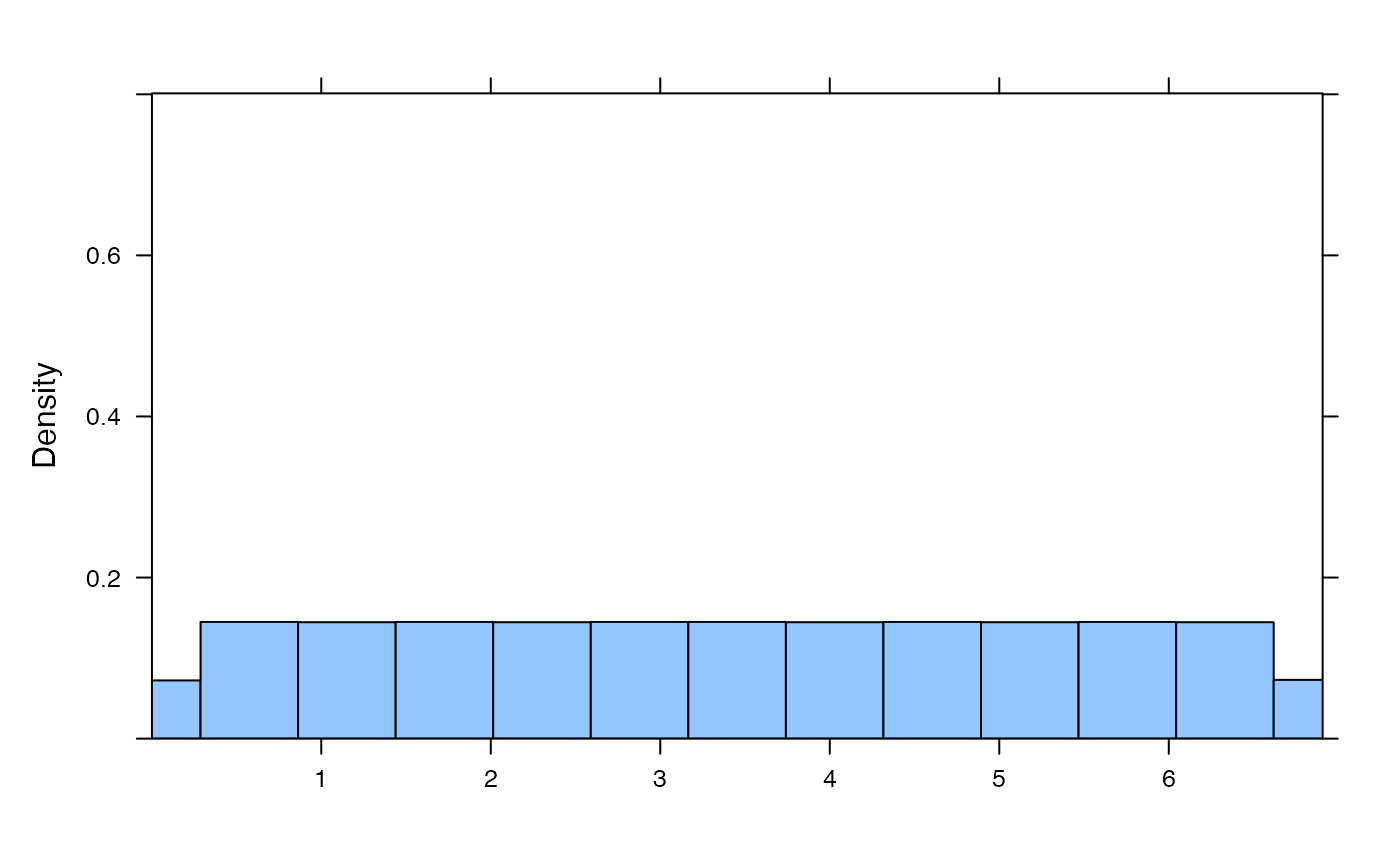 plotDist('binom', params=list( 25, .25)) # explicit params
plotDist('binom', params=list( 25, .25)) # explicit params
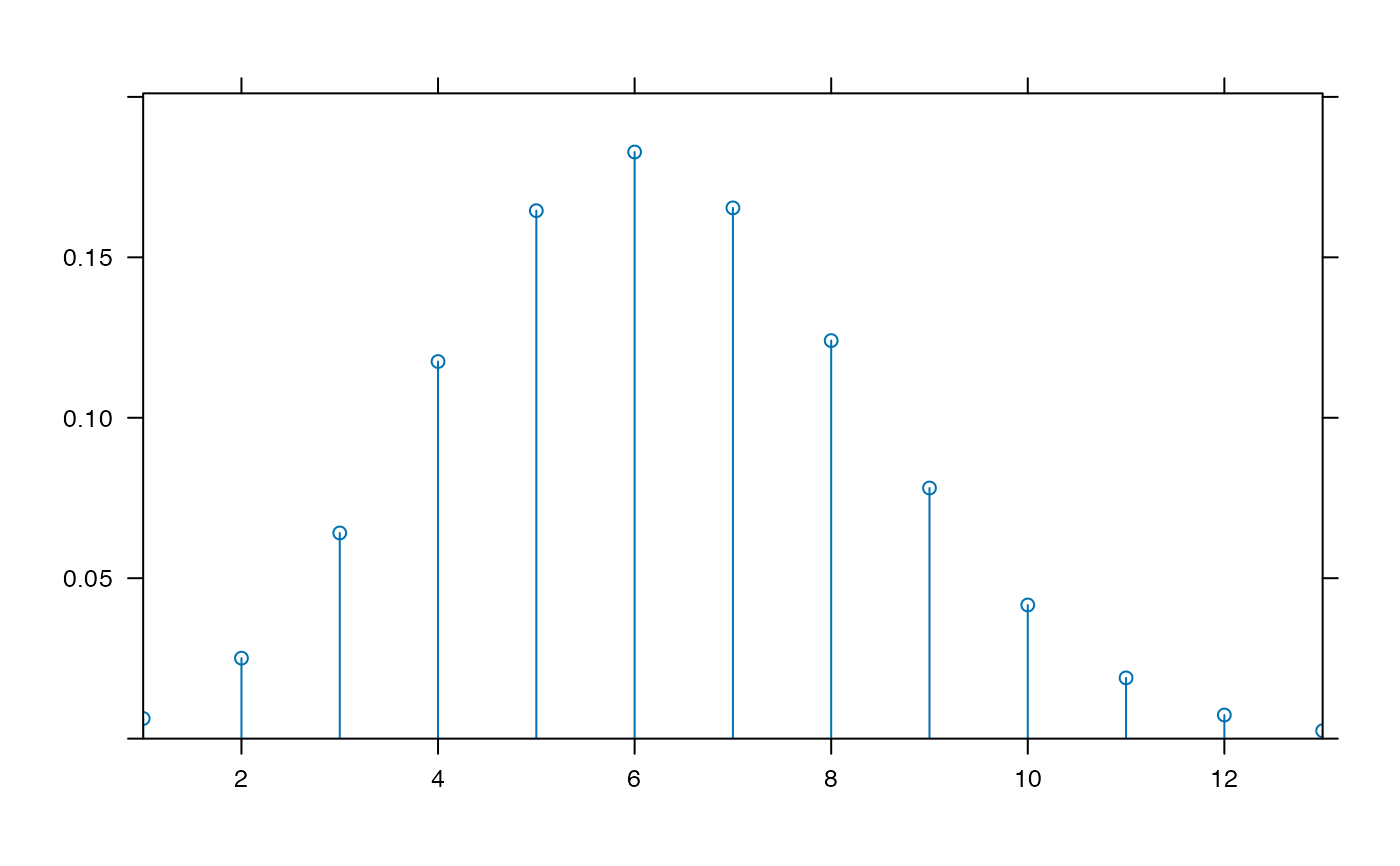 plotDist('binom', 25, .25) # params inferred
plotDist('binom', 25, .25) # params inferred
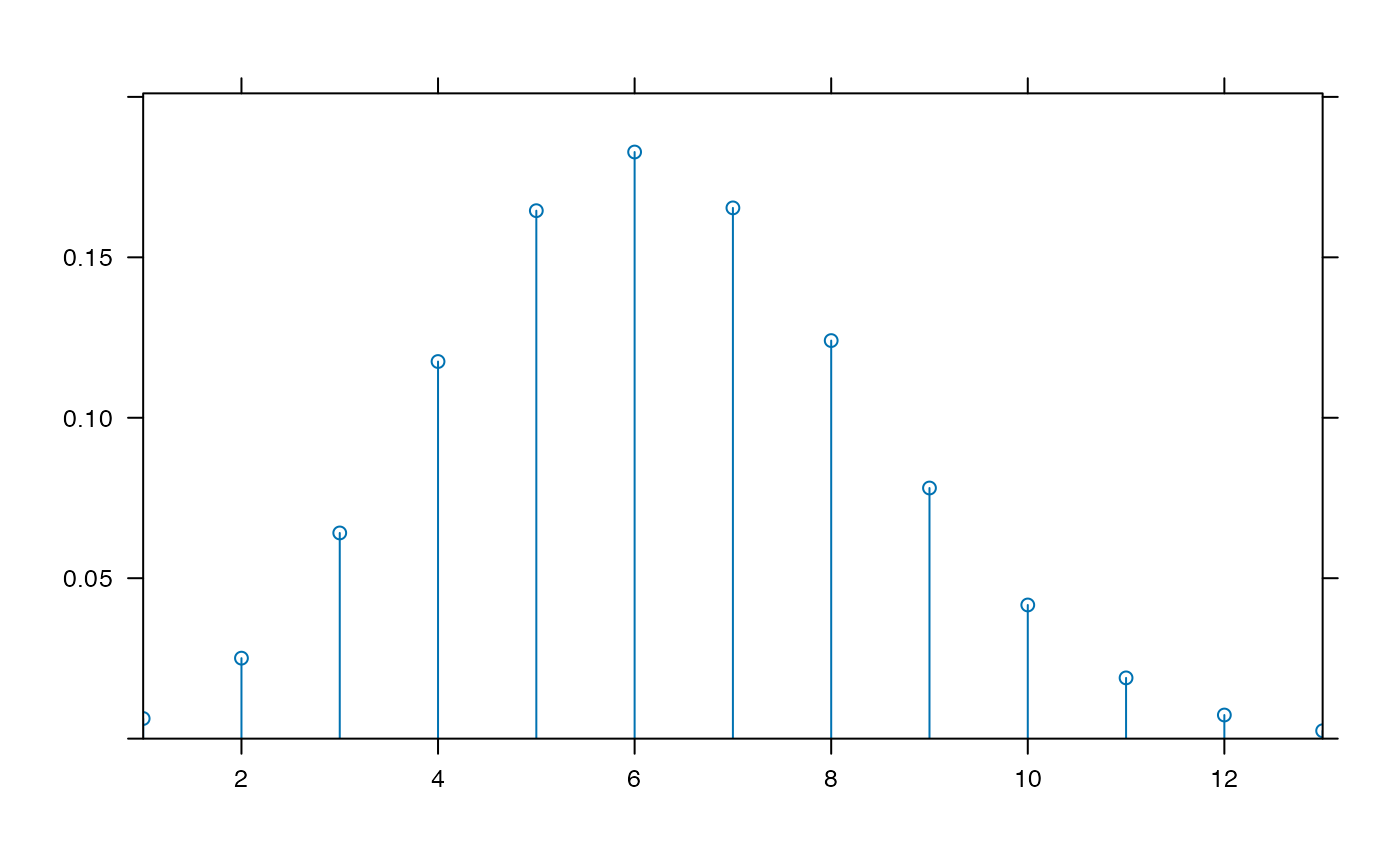 plotDist('norm', mean=100, sd=10, kind='cdf') # params inferred
plotDist('norm', mean=100, sd=10, kind='cdf') # params inferred
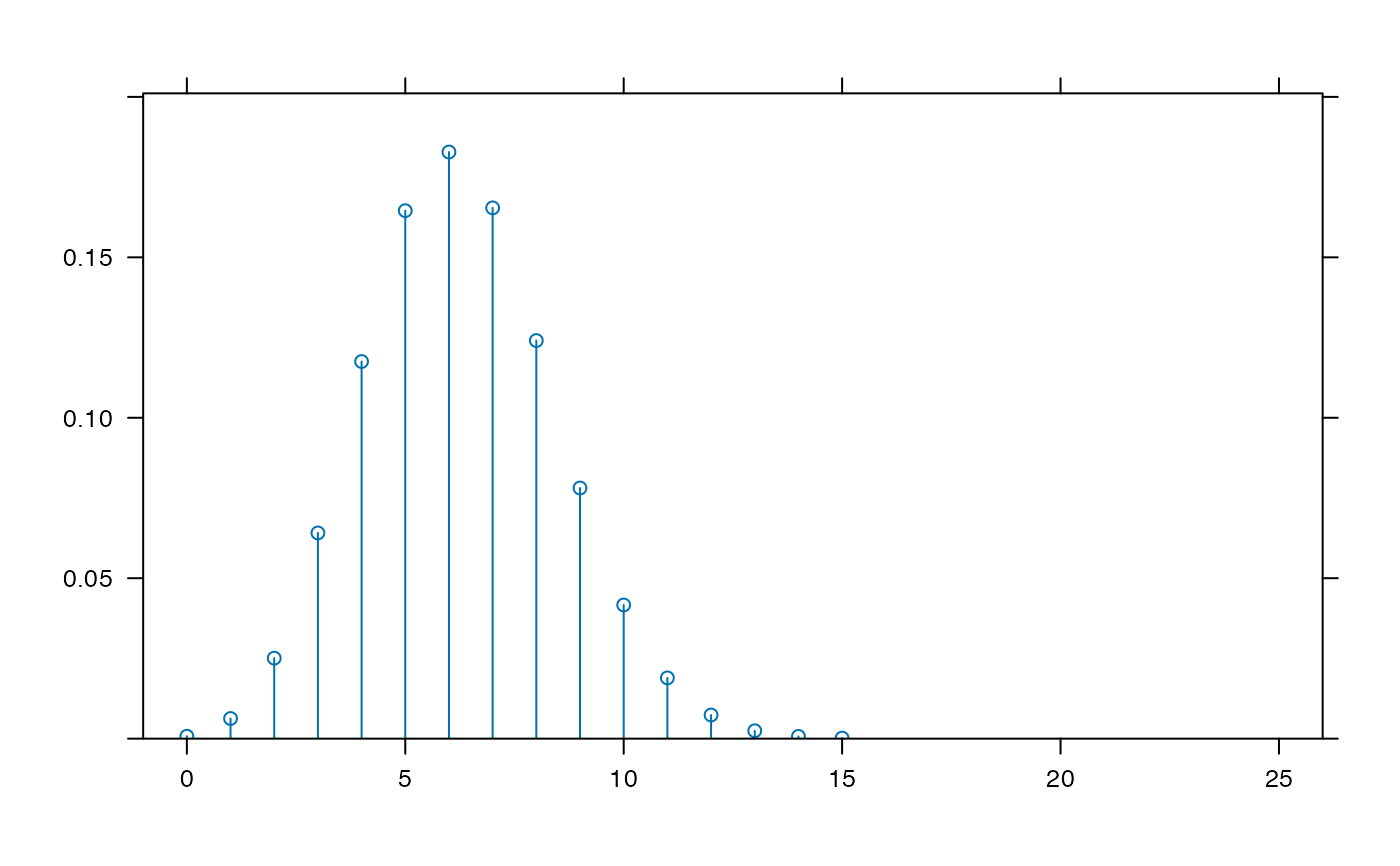
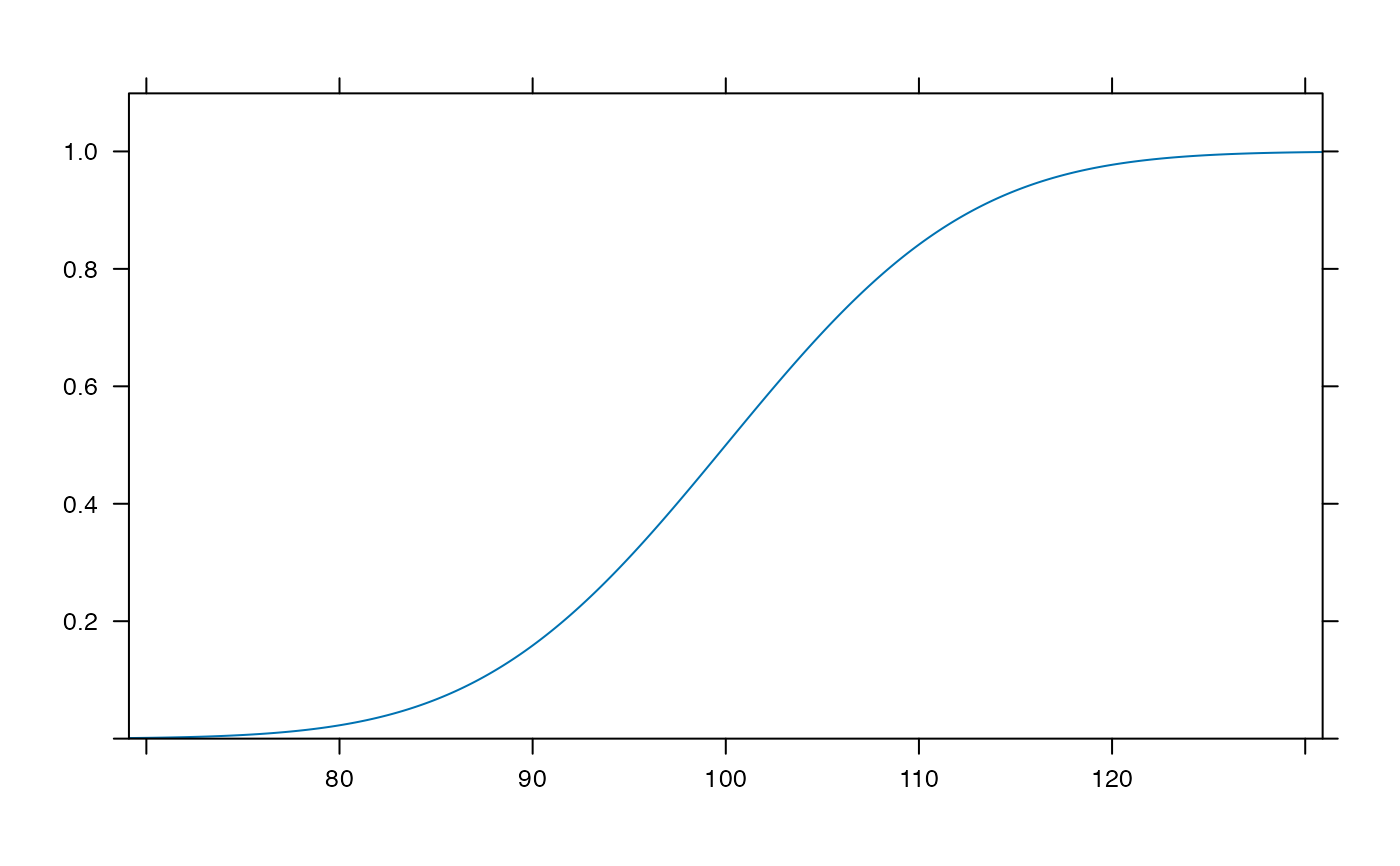 plotDist('binom', 25, .25, xlim=c(-1,26) ) # params inferred
plotDist('binom', 25, .25, xlim=c(-1,26) ) # params inferred
 plotDist('binom', params=list( 25, .25), kind='cdf')
plotDist('binom', params=list( 25, .25), kind='cdf')
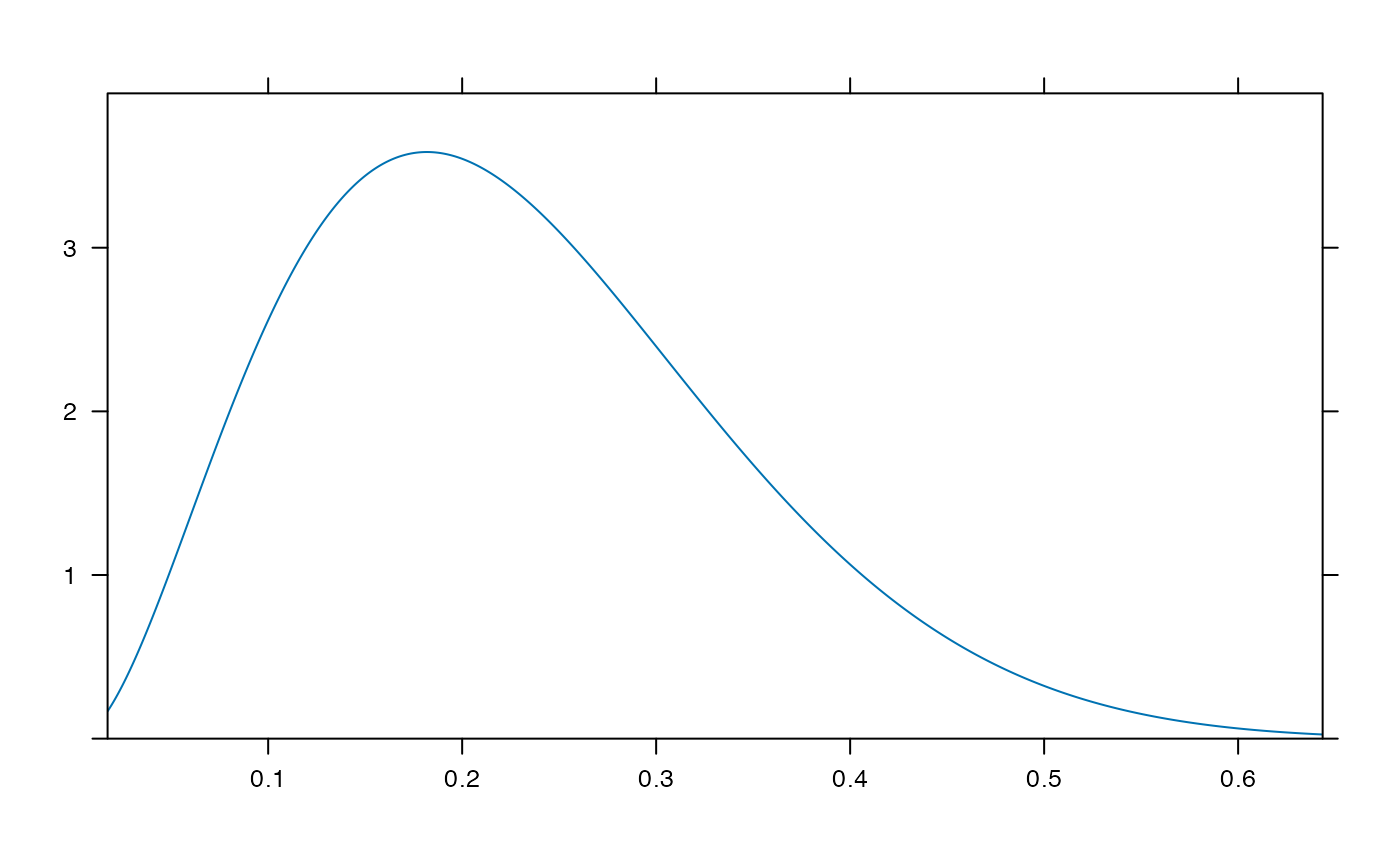
 plotDist('beta', params=list( 3, 10), kind='density')
plotDist('beta', params=list( 3, 10), kind='density')
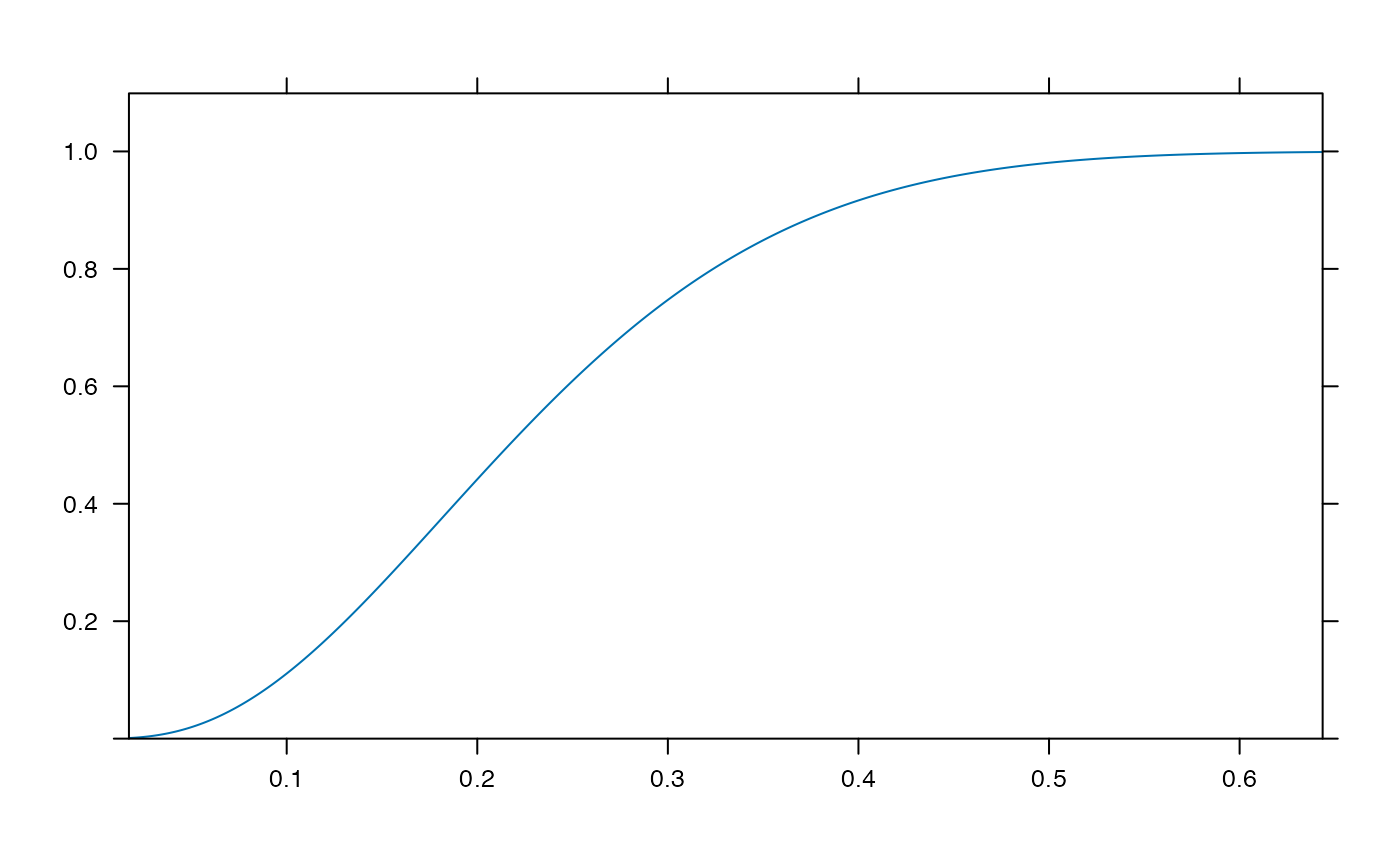
 plotDist('beta', params=list( 3, 10), kind='cdf')
plotDist('beta', params=list( 3, 10), kind='cdf')
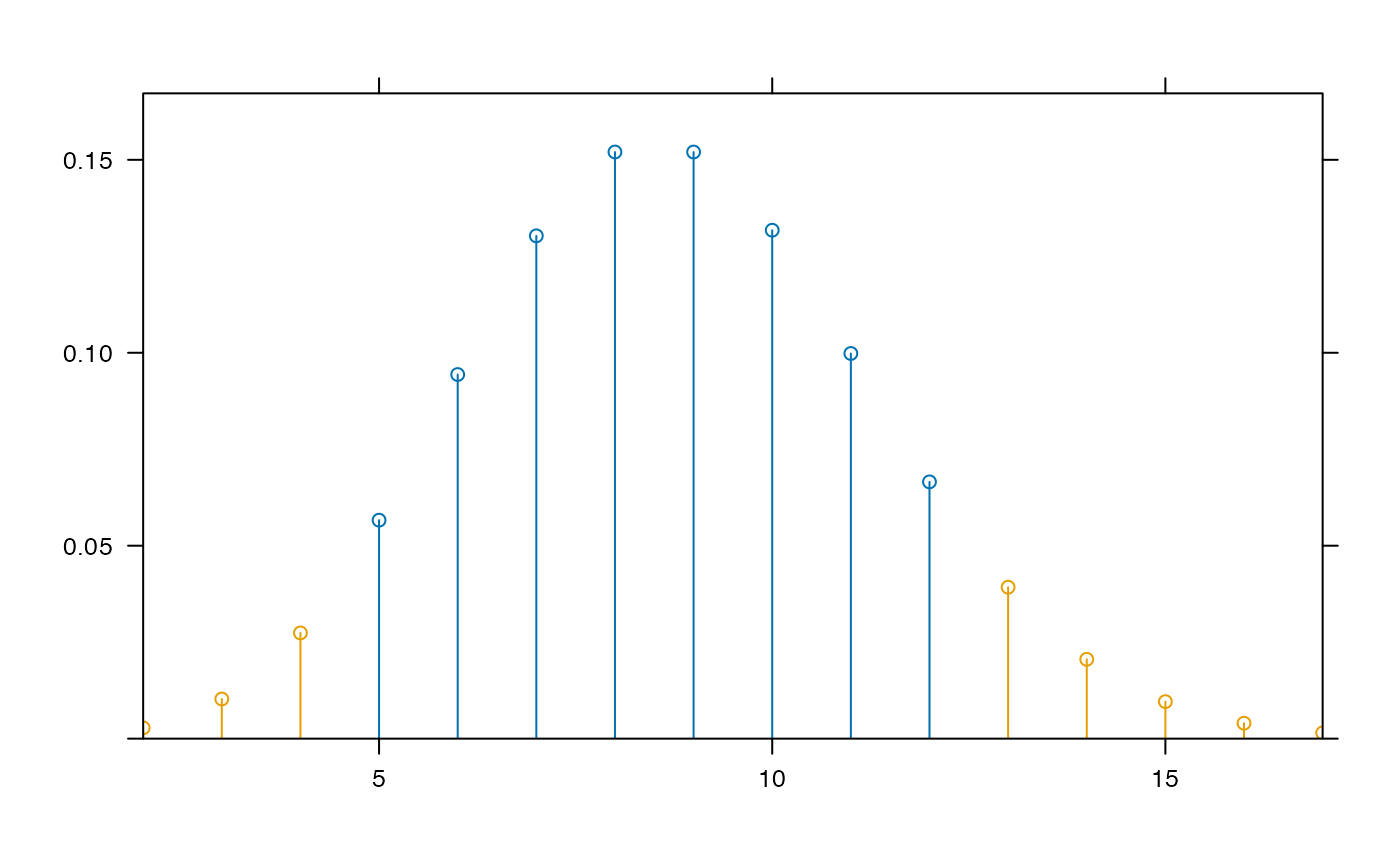
 plotDist( "binom", params=list(35,.25),
groups= y < dbinom(qbinom(0.05, 35, .25), 35,.25) )
plotDist( "binom", params=list(35,.25),
groups= y < dbinom(qbinom(0.05, 35, .25), 35,.25) )
 plotDist( "binom", params=list(35,.25),
groups= y < dbinom(qbinom(0.05, 35, .25), 35,.25),
kind='hist')
plotDist( "binom", params=list(35,.25),
groups= y < dbinom(qbinom(0.05, 35, .25), 35,.25),
kind='hist')
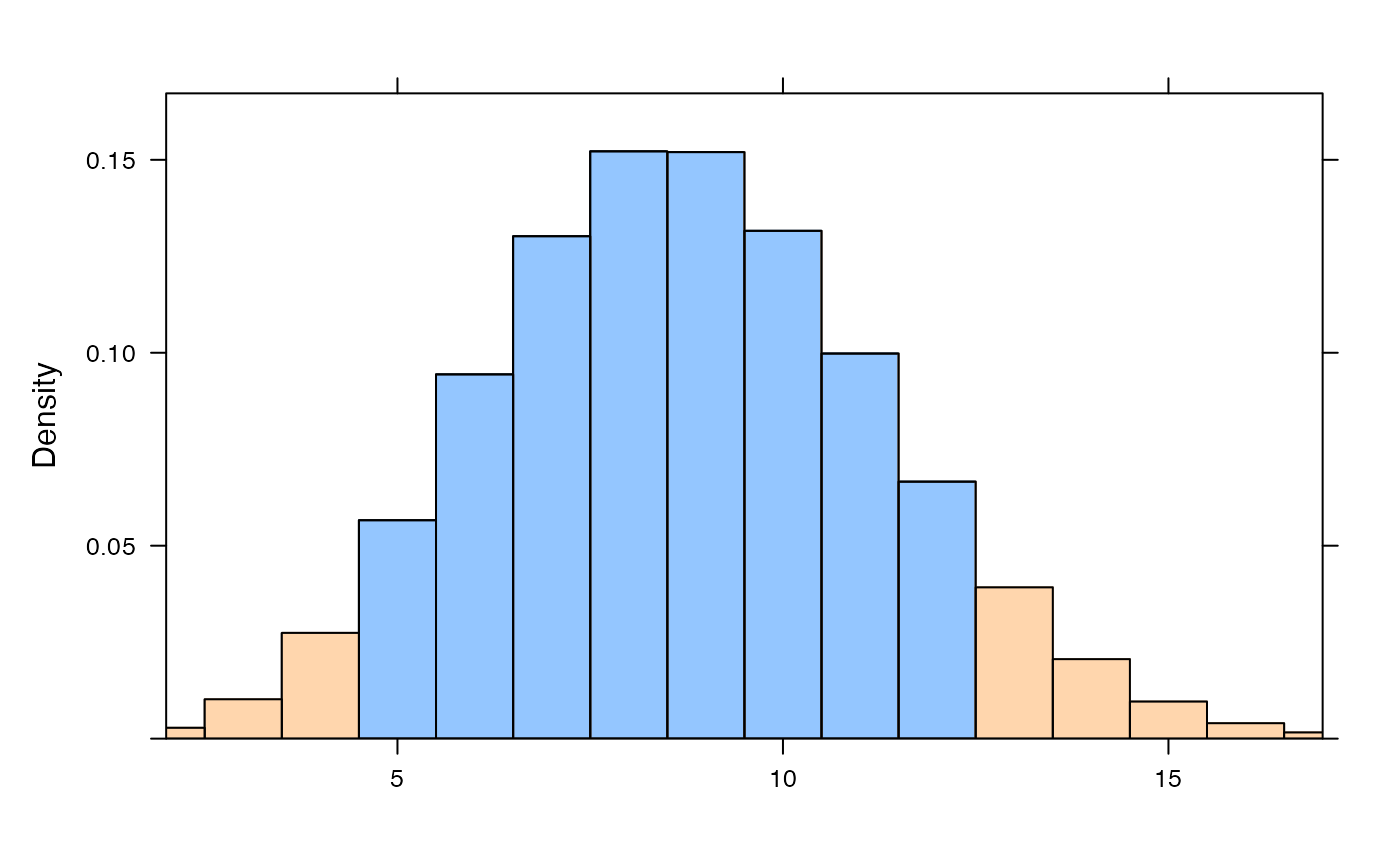 plotDist("norm", mean=10, sd=2, col="blue", type="h")
plotDist("norm", mean=10, sd=2, col="blue", type="h")
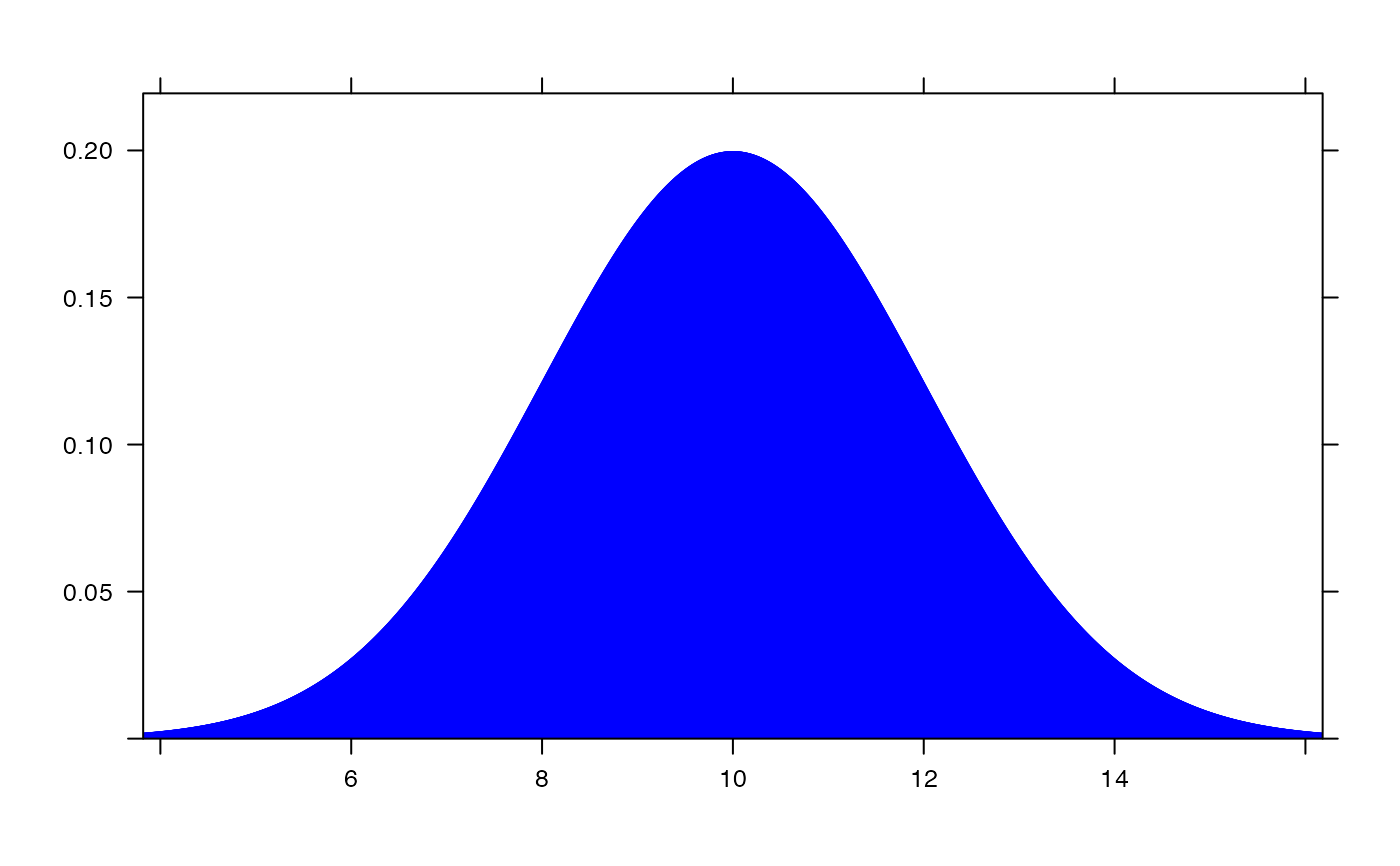 plotDist("norm", mean=12, sd=2, col="red", type="h", under=TRUE)
plotDist("norm", mean=12, sd=2, col="red", type="h", under=TRUE)
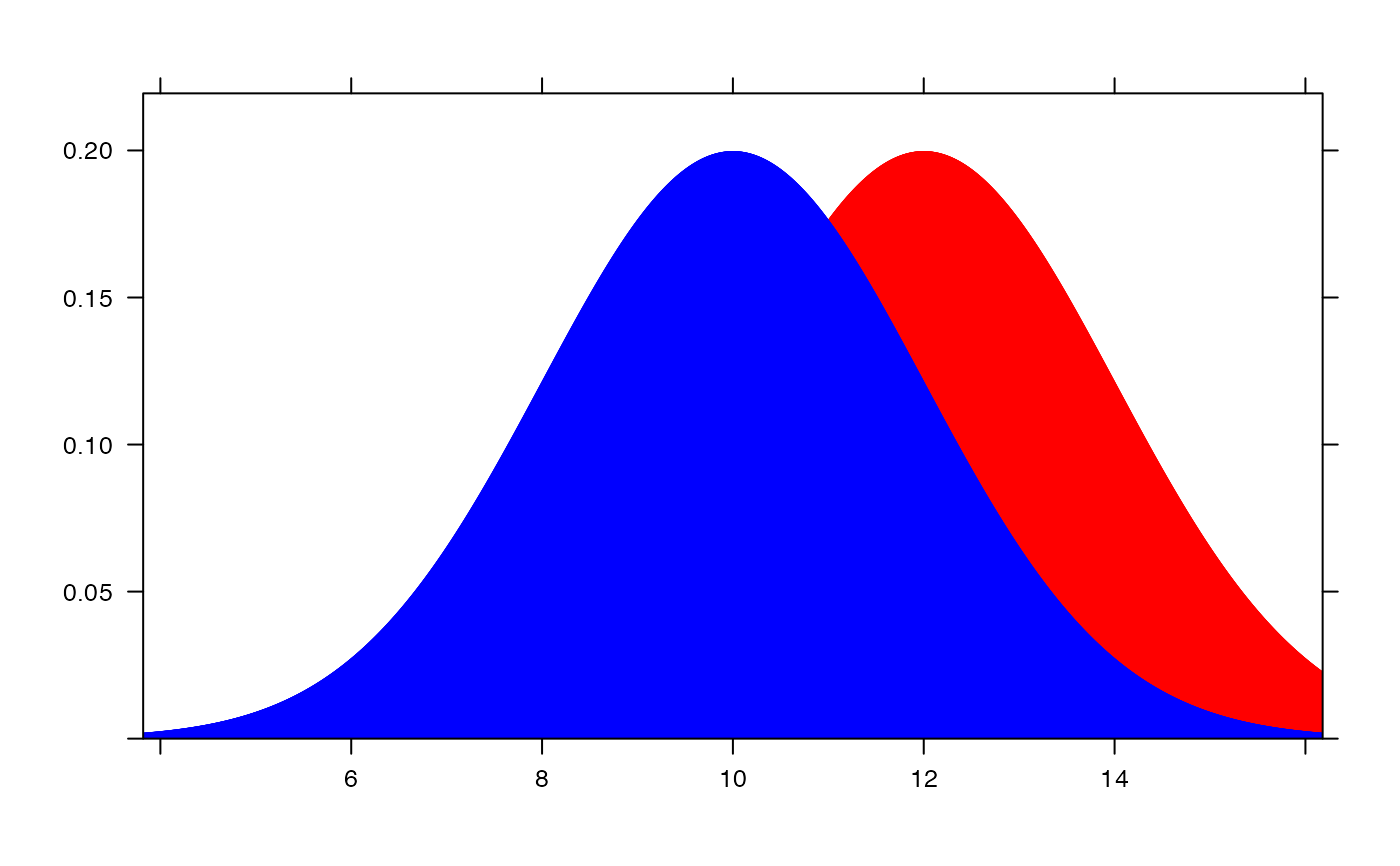 plotDist("binom", size=100, prob=.30) +
plotDist("norm", mean=30, sd=sqrt(100 * .3 * .7))
plotDist("binom", size=100, prob=.30) +
plotDist("norm", mean=30, sd=sqrt(100 * .3 * .7))
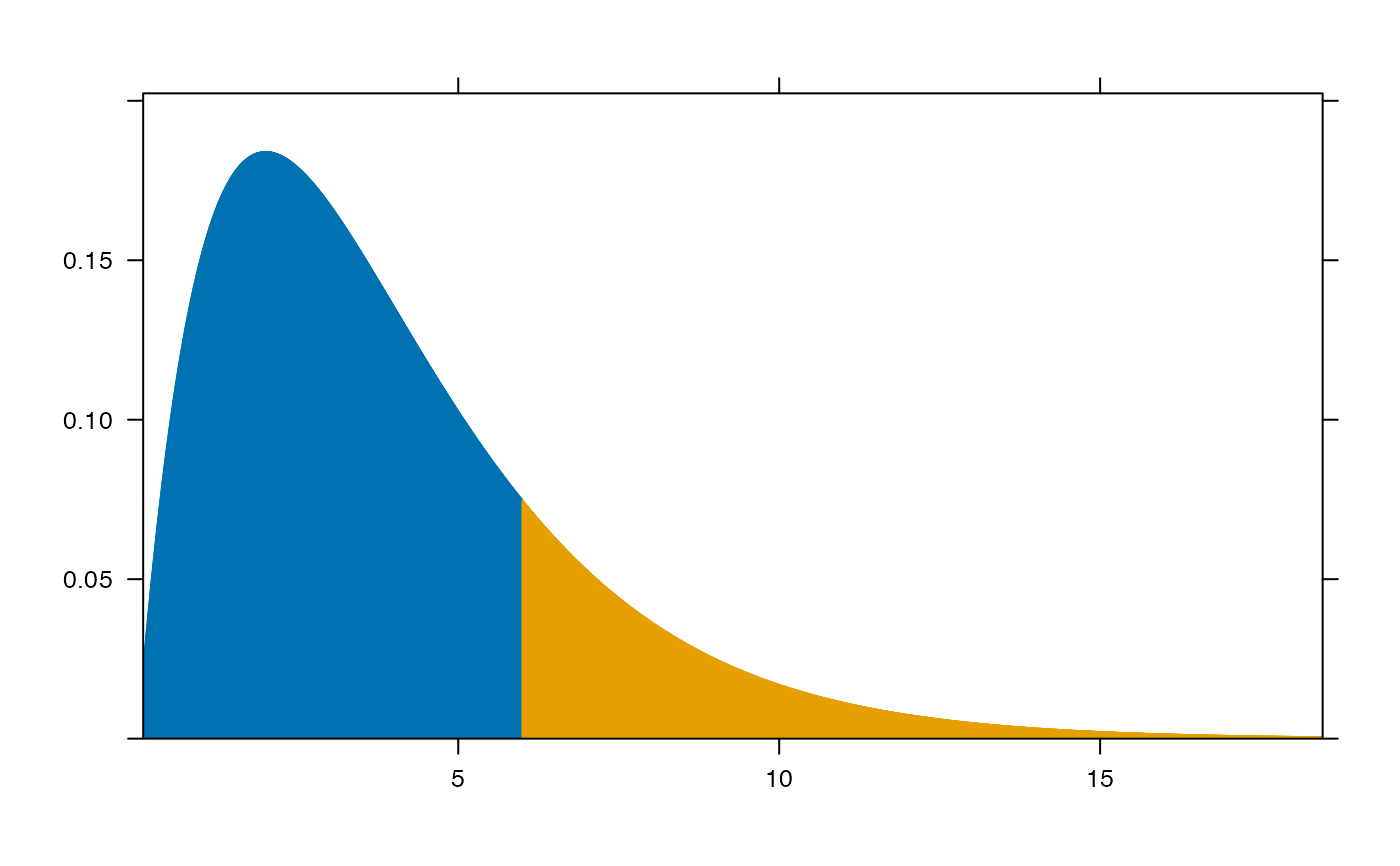
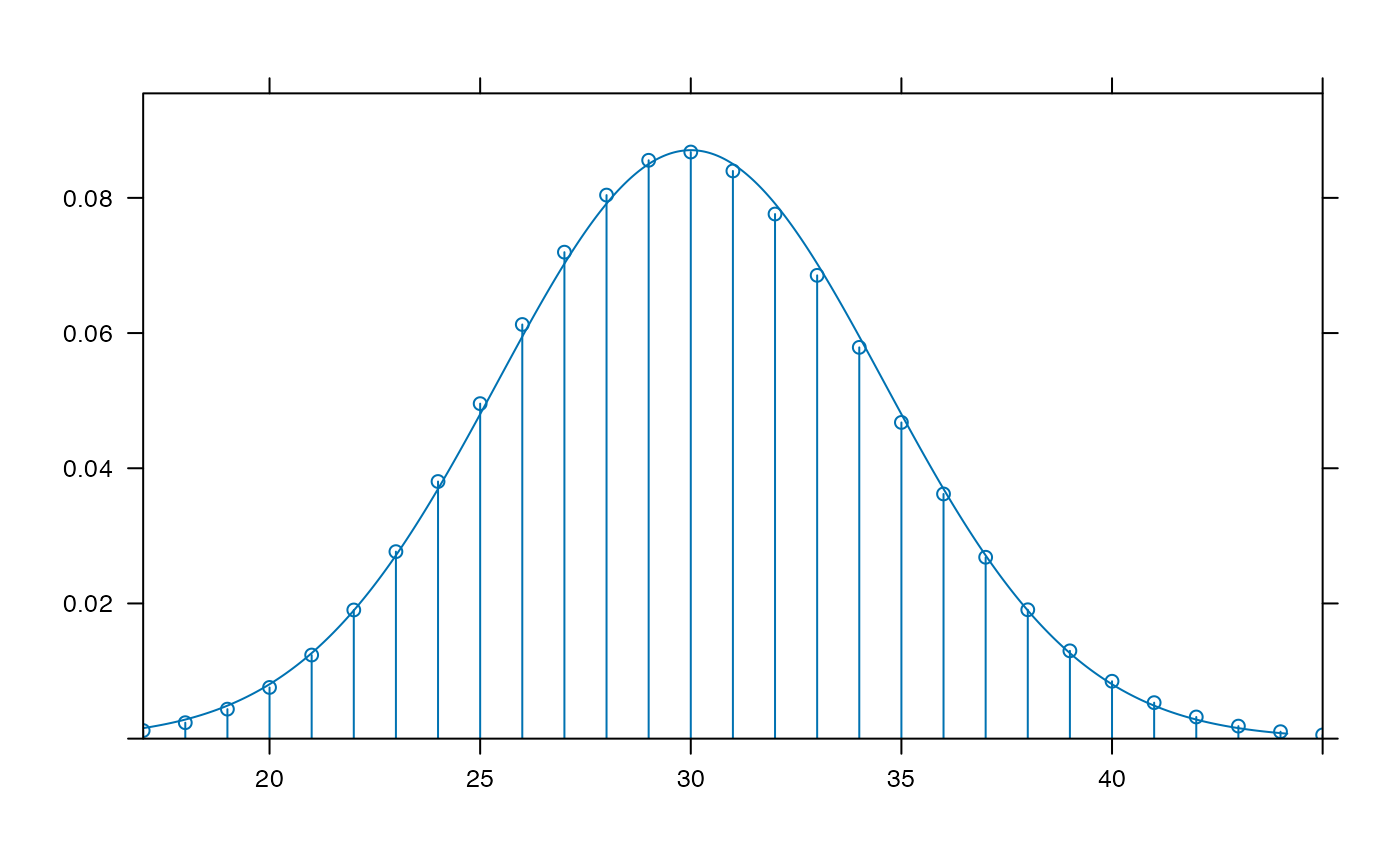 plotDist("chisq", df=4, groups = x > 6, type="h")
plotDist("chisq", df=4, groups = x > 6, type="h")
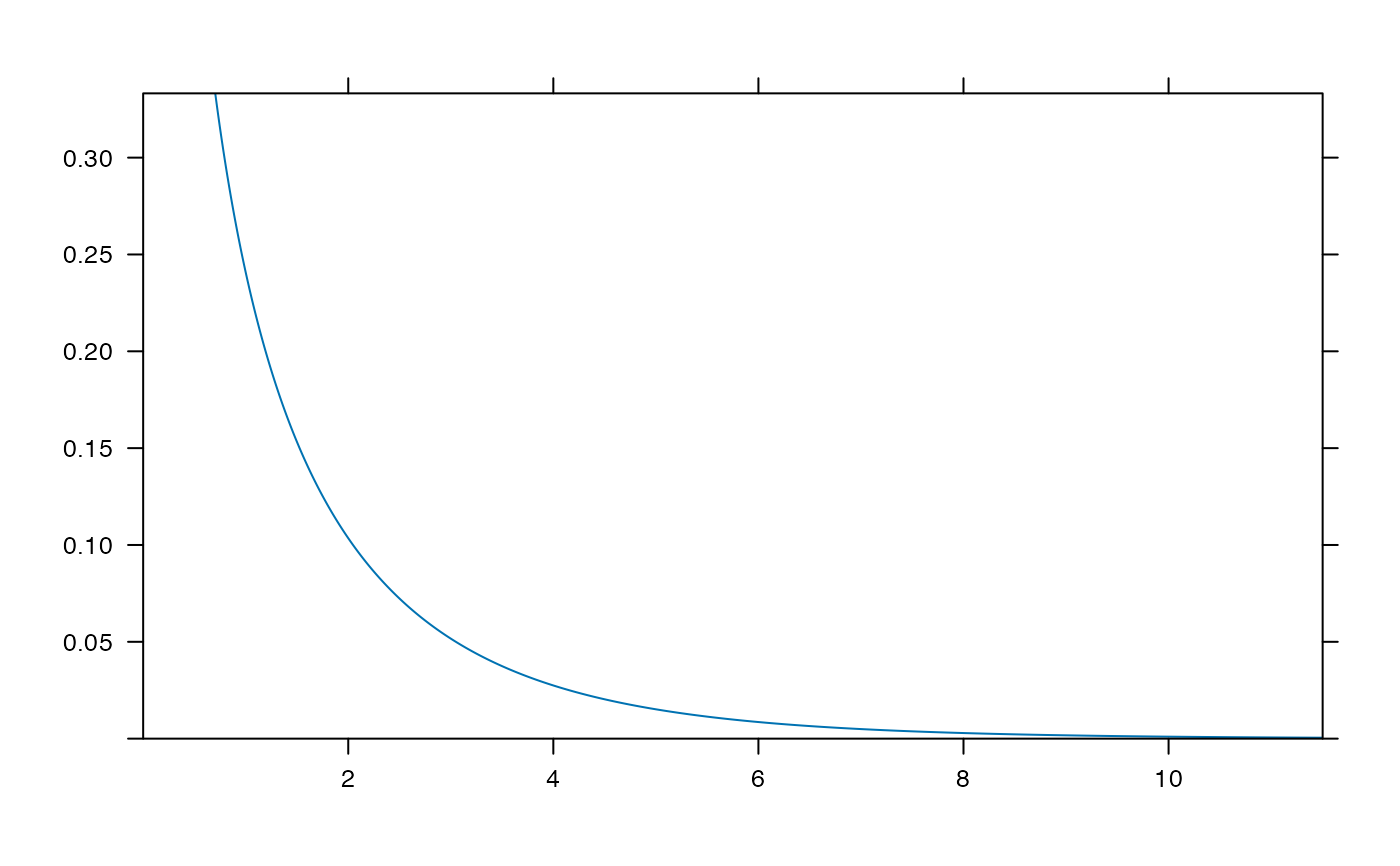
 plotDist("f", df1=1, df2 = 99)
plotDist("f", df1=1, df2 = 99)
 if (require(mosaicData)) {
histogram( ~age|sex, data=HELPrct)
m <- mean( ~age|sex, data=HELPrct)
s <- sd(~age|sex, data=HELPrct)
plotDist( "norm", mean=m[1], sd=s[1], col="red", add=TRUE, packets=1)
plotDist( "norm", mean=m[2], sd=s[2], col="blue", under=TRUE, packets=2)
}
if (require(mosaicData)) {
histogram( ~age|sex, data=HELPrct)
m <- mean( ~age|sex, data=HELPrct)
s <- sd(~age|sex, data=HELPrct)
plotDist( "norm", mean=m[1], sd=s[1], col="red", add=TRUE, packets=1)
plotDist( "norm", mean=m[2], sd=s[2], col="blue", under=TRUE, packets=2)
}